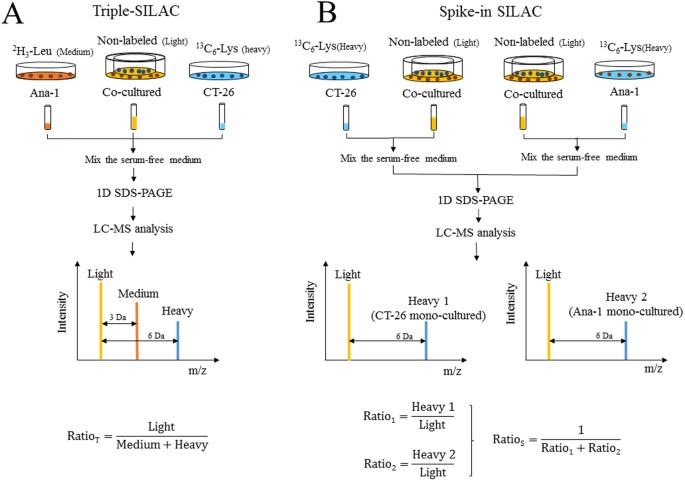
IJMS | Free Full-Text | SILAC-Based Quantification of TGFBR2-Regulated Protein Expression in Extracellular Vesicles of Microsatellite Unstable Colorectal Cancers | HTML

A practical recipe for stable isotope labeling by amino acids in cell culture (SILAC) | Semantic Scholar

Encoding quantitative information into whole proteomes with SILAC.Cells... | Download Scientific Diagram

Light or heavy isotope labeled amino acids (Lys, +6 Da; Arg, +10 Da)... | Download Scientific Diagram

Light or heavy isotope labeled amino acids (Lys, +6 Da; Arg, +10 Da)... | Download Scientific Diagram

Light or heavy isotope labeled amino acids (Lys, +6 Da; Arg, +10 Da)... | Download Scientific Diagram
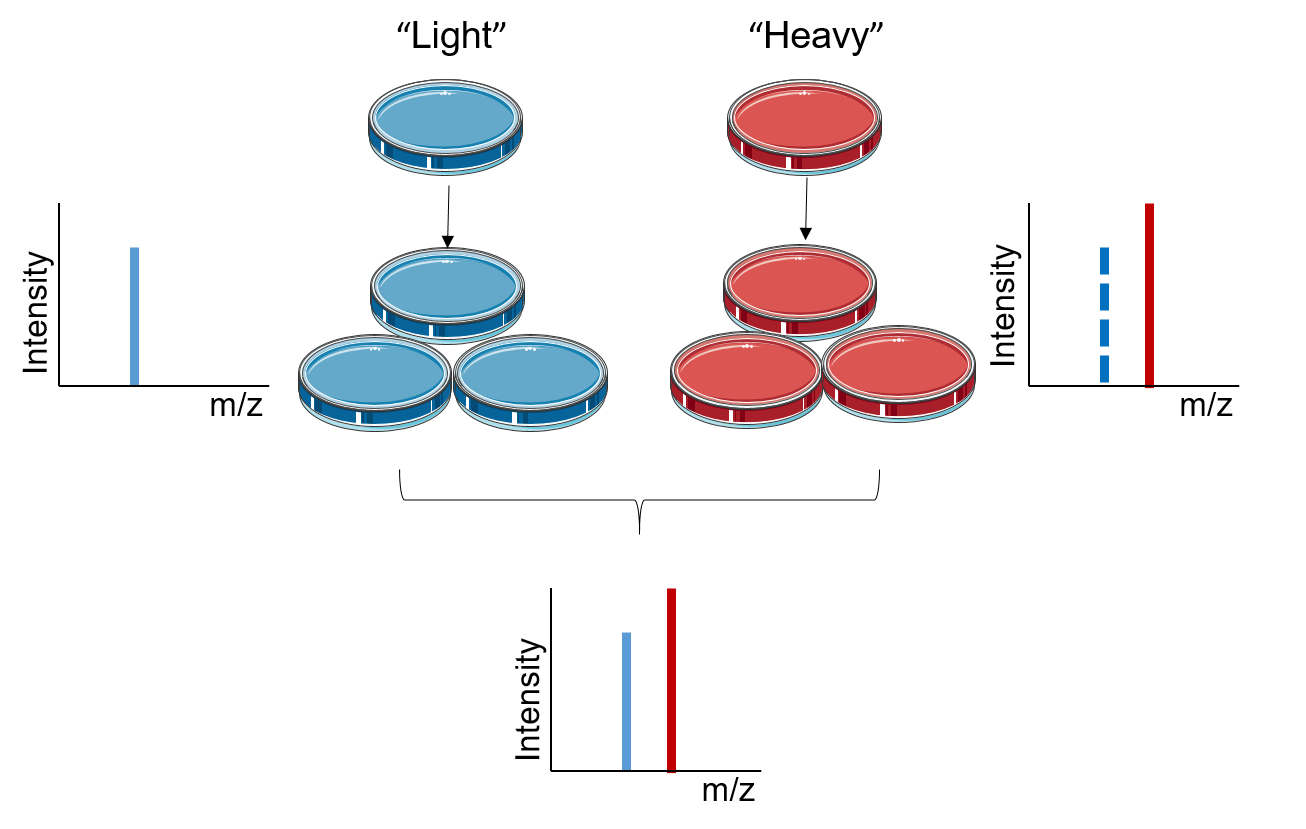
Stable Isotope Labeling Using Amino Acids in Cell Culture (SILAC): Principles, Workflow, and Applications – Creative Proteomics Blog

Amino acid based labeling techniques. (A) SILAC labeling. Proteins are... | Download Scientific Diagram

Use of stable isotope labeling by amino acids in cell culture as a spike-in standard in quantitative proteomics | Nature Protocols
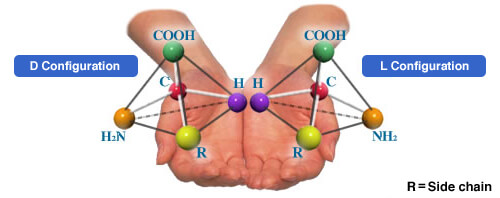
What are L-, D-, and DL-Amino Acids? | Enhancing Life with Amino Acids | About us | Ajinomoto Group Global Website - Eat Well, Live Well.

Schematic representation of a stable isotope labelling with amino acids... | Download Scientific Diagram

Comparison of Isotope-labeled Amino Acid Incorporation Rates (CILAIR) Provides a Quantitative Method to Study Tissue Secretomes - Molecular & Cellular Proteomics

Intercellular Transfer of Proteins as Identified by Stable Isotope Labeling of Amino Acids in Cell Culture* - Journal of Biological Chemistry

Quantitative, Time-Resolved Proteomic Analysis by Combining Bioorthogonal Noncanonical Amino Acid Tagging and Pulsed Stable Isotope Labeling by Amino Acids in Cell Culture* - Molecular & Cellular Proteomics